dat <- lavaan::HolzingerSwineford1939
mod <- "
visual =~ x1 + x2 + x3
textual =~ x4 + x5 + x6
speed =~ x7 + x8 + x9
"
fit <- acfa(mod, dat, verbose = FALSE)Purpose
After fitting a Bayesian SEM with INLAvaan, we often want to draw samples from the model itself—either using posterior or prior parameter values. The sampling() function does exactly this: it propagates parameter draws through the full generative chain
producing samples that are not tied to any individual observation. This is distinct from predict(), which returns individual-specific factor scores .
Typical use cases include posterior predictive checks (PPCs) and prior predictive checks.
The generative model
Let () denote one parameter draw. From this draw the SEM matrices , , , , , and are constructed. The generative chain is:
1. Latent variables.
2. Observed variables.
When prior = FALSE (default), comes from the posterior; when prior = TRUE, each parameter is drawn independently from its prior.
Quick start
Parameter samples
The default type "lavaan" returns an matrix of lavaan-side (constrained) parameter draws:
theta_post <- sampling(fit, type = "lavaan", nsamp = 2000)
dim(theta_post)
#> [1] 2000 21
head(theta_post[, 1:4])
#> visual=~x2 visual=~x3 textual=~x5 textual=~x6
#> [1,] 0.6216553 0.9058102 1.124324 0.9962591
#> [2,] 0.5512186 0.8048597 1.213441 0.9163449
#> [3,] 0.5589617 0.7005065 0.980363 0.9248275
#> [4,] 0.5567215 0.7637923 1.141509 1.0359674
#> [5,] 0.5948079 1.0228482 1.073171 0.8665535
#> [6,] 0.3633649 0.7067464 1.089029 0.8697055Latent and observed samples
Everything at once
Comparing samples to the fitted marginals
INLAvaan approximates each marginal posterior with a skew-normal density. We can overlay the sampling() histogram (drawn via the copula method) on top of the fitted density curve stored in pdf_data to verify that they agree.
# 1. Draw copula samples
samp_cop <- sampling(fit, type = "lavaan", nsamp = 1000, samp_copula = TRUE)
# 2. Retrieve the fitted skew-normal densities
int <- fit@external$inlavaan_internal
pdf_data <- int$pdf_data
# 3. Build a long data frame for ggplot
par_names <- colnames(samp_cop)
hist_df <- data.frame(
param = rep(par_names, each = nrow(samp_cop)),
value = as.vector(samp_cop)
)
hist_df$param <- factor(hist_df$param, levels = par_names)
curve_df <- do.call(rbind, Map(function(nm, df) {
data.frame(param = nm, x = df$x, y = df$y)
}, par_names, pdf_data[par_names]))
curve_df$param <- factor(curve_df$param, levels = par_names)
# 4. Plot
ggplot(hist_df, aes(x = value)) +
geom_histogram(aes(y = after_stat(density)), bins = 50,
fill = "grey40", alpha = 0.5) +
geom_line(data = curve_df, aes(x = x, y = y),
colour = "#00A6AA", linewidth = 0.8) +
facet_wrap(~param, scales = "free") +
labs(x = "Parameter value", y = "Density",
title = "Copula samples vs. fitted skew-normal posterior") +
theme_minimal(base_size = 11)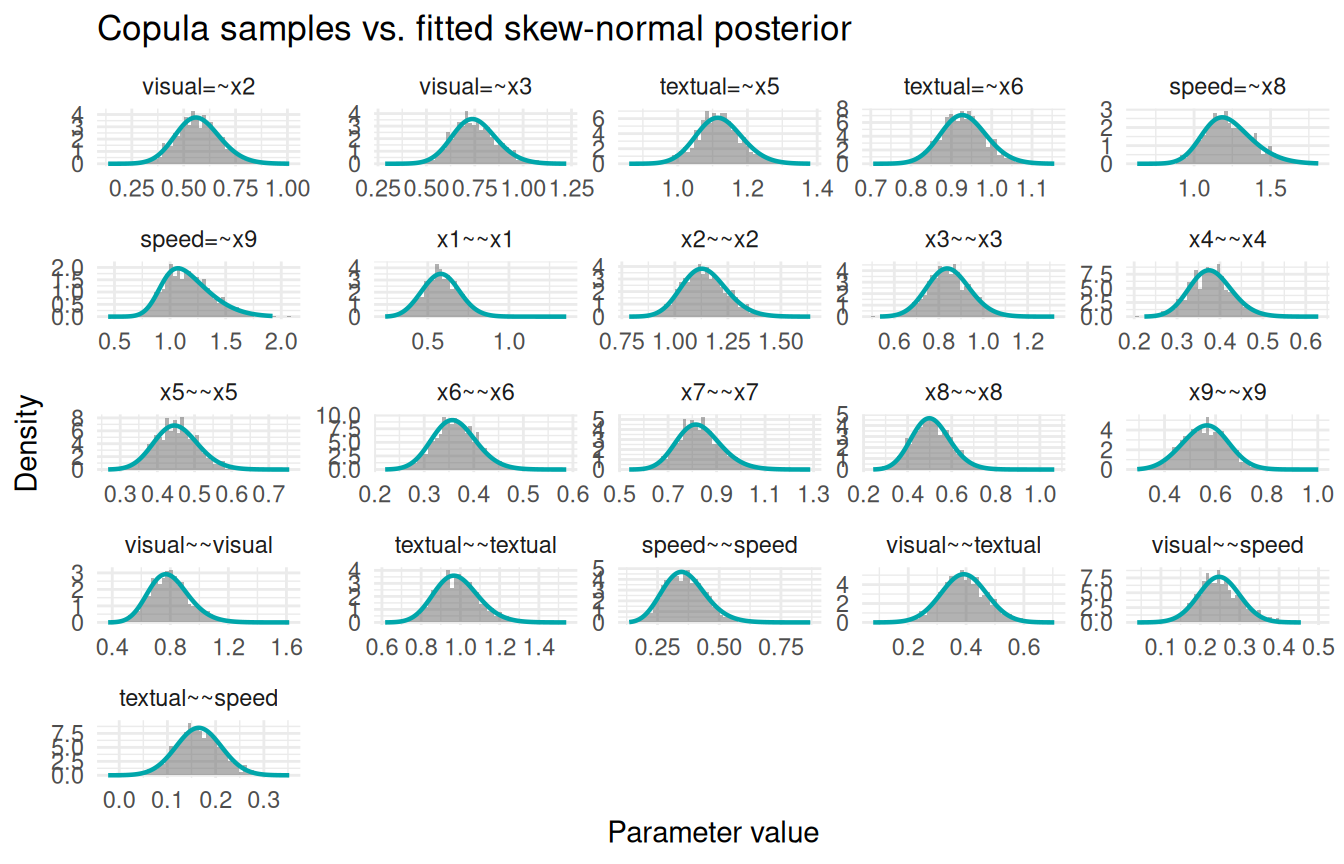
Copula vs. Gaussian sampling
By default, sampling() uses the copula method, which respects the skew-normal marginals. Setting samp_copula = FALSE uses the multivariate Gaussian (Laplace) approximation instead. The difference is most visible for parameters with asymmetric posteriors, such as variance components.
samp_gauss <- sampling(fit, type = "lavaan", nsamp = 5000, samp_copula = FALSE)
cop_df <- data.frame(
param = rep(par_names, each = nrow(samp_cop)),
value = as.vector(samp_cop),
method = "Copula"
)
gauss_df <- data.frame(
param = rep(par_names, each = nrow(samp_gauss)),
value = as.vector(samp_gauss),
method = "Gaussian"
)
both_df <- rbind(cop_df, gauss_df)
both_df$param <- factor(both_df$param, levels = par_names)
both_df$method <- factor(both_df$method, levels = c("Copula", "Gaussian"))
ggplot(both_df, aes(x = value, fill = method)) +
geom_histogram(aes(y = after_stat(density)), bins = 50,
alpha = 0.45, position = "identity") +
facet_wrap(~param, scales = "free") +
scale_fill_manual(values = c(Copula = "#00A6AA", Gaussian = "#F18F00")) +
labs(x = "Parameter value", y = "Density", fill = "Method",
title = "Copula vs. Gaussian posterior sampling") +
theme_minimal(base_size = 11) +
theme(legend.position = "top")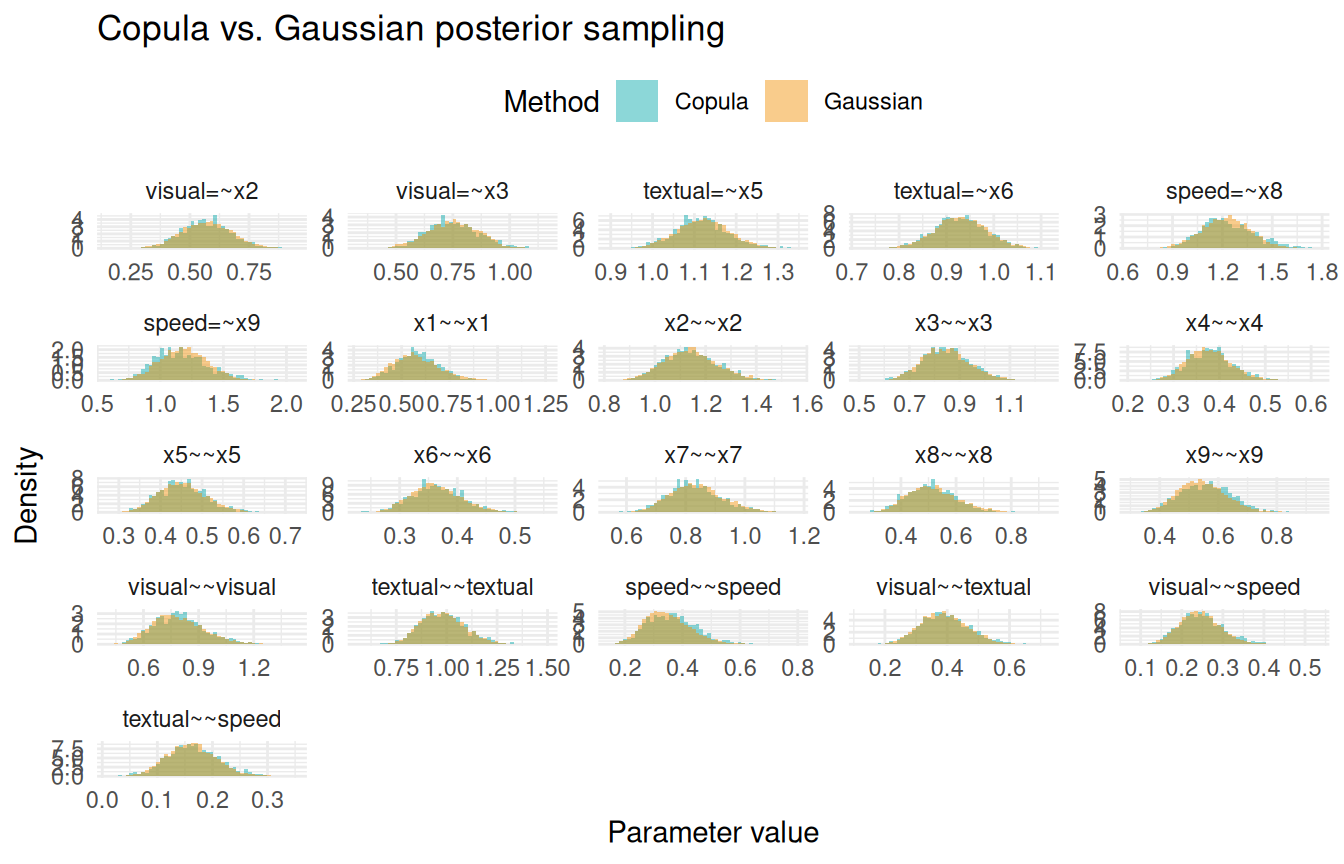
For symmetric posteriors (e.g., factor loadings), the two methods are nearly identical. For positively-skewed posteriors (e.g., residual variances), the copula method captures the asymmetry while the Gaussian approximation is symmetric by construction.
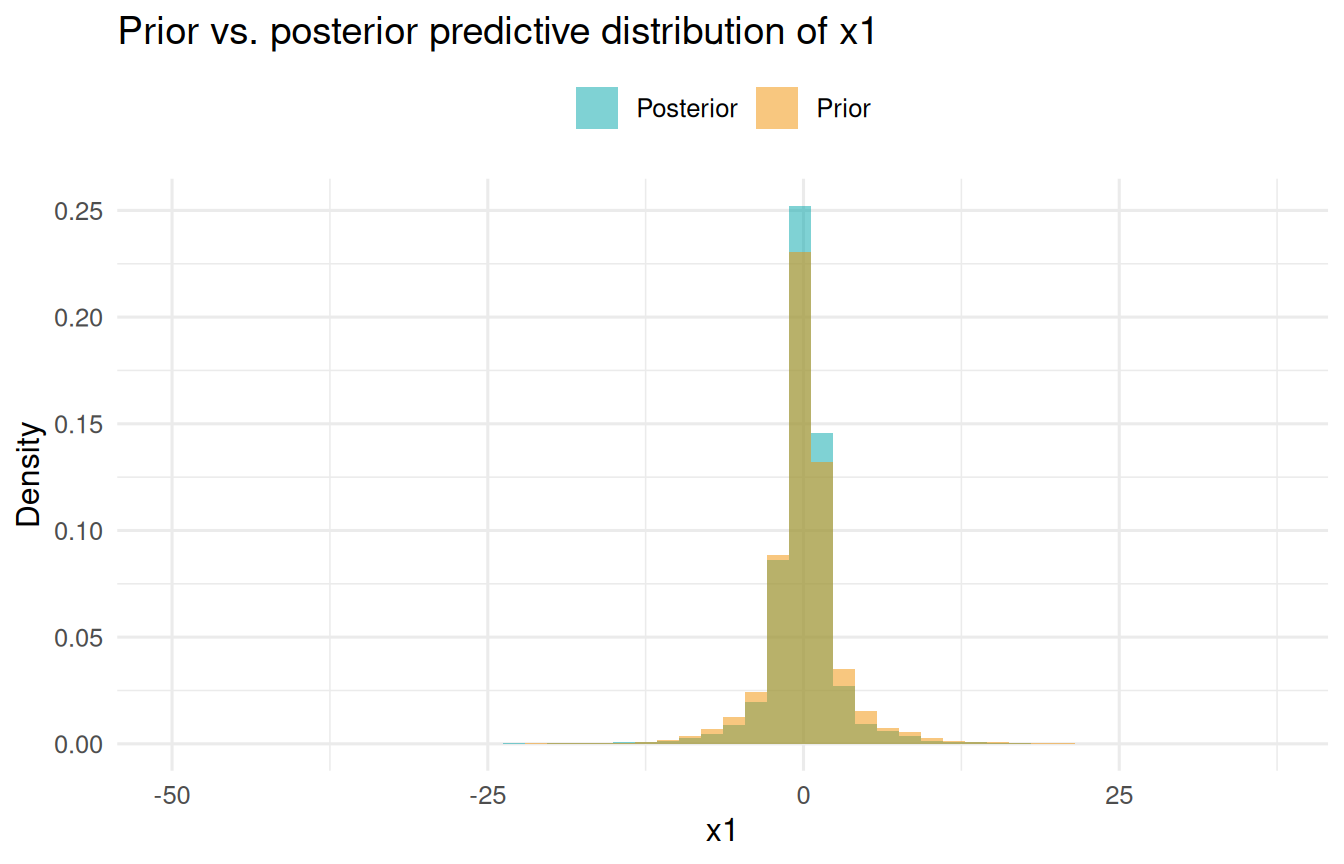
Prior predictive sampling
Setting prior = TRUE draws parameters from the prior distribution (independently, ignoring the data) and propagates them through the generative model. This is useful for checking whether the priors imply sensible ranges for the observed data.
df_pp <- data.frame(
x1 = c(y_prior[, "x1"], y_post[, "x1"]),
source = rep(c("Prior", "Posterior"), each = 2000)
)
ggplot(df_pp, aes(x = x1, fill = source)) +
geom_histogram(aes(y = after_stat(density)), bins = 50,
alpha = 0.5, position = "identity") +
scale_fill_manual(values = c(Prior = "#F18F00", Posterior = "#00A6AA")) +
labs(x = "x1", y = "Density", fill = NULL,
title = "Prior vs. posterior predictive distribution of x1") +
theme_minimal(base_size = 12) +
theme(legend.position = "top")