HS.model <- "
visual =~ x1 + x2 + x3
textual =~ x4 + x5 + x6
speed =~ x7 + x8 + x9
"
fit <- acfa(HS.model, HolzingerSwineford1939, verbose = FALSE)Introduction
Classical SEM fit indices (RMSEA, CFI, TLI, etc.) are computed from a single maximum-likelihood chi-square statistic. In Bayesian SEM, the parameters are random, so each posterior draw yields a different chi-square value and, therefore, a different set of fit indices. INLAvaan provides Bayesian analogues of the most common indices, following the framework of Garnier-Villarreal and Jorgensen (2020) (as implemented in blavaan).
This vignette covers:
- Mathematical definitions of the Bayesian fit indices.
- The two rescaling methods (
"devM"and"MCMC"). - A worked example with the Holzinger–Swineford data.
- Comparison with a baseline model (incremental indices).
- Differences between INLAvaan and blavaan.
Mathematical background
Deviance and chi-square
Let denote the -th posterior draw of the model parameters (). The per-sample deviance chi-square is where is the log-likelihood of the saturated model (sample moments equal model moments) and is the log-likelihood evaluated at the -th draw. This equals , where is the ML discrepancy function.
Rescaling
INLAvaan supports two rescaling methods, controlled by the rescale argument:
"devM" (default). Uses the DIC-based effective number of parameters . If is unreasonable ( or ), INLAvaan falls back to using (the number of free parameters).
"MCMC". Uses the classical chi-square with scaling. where is the number of free parameters and is the number of groups.
In both cases the per-sample noncentrality parameter is .
Absolute fit indices
The following indices are computed at each posterior draw , generating a posterior distribution.
BRMSEA (Bayesian RMSEA).
BGammaHat. where is the total number of observed variables across groups.
Adjusted BGammaHat.
BMc (McDonald’s centrality index).
Incremental fit indices
Incremental indices compare the target model against a baseline (null) model. Let , , and denote the corresponding quantities for the baseline model.
BCFI (Bayesian CFI).
BTLI (Bayesian TLI).
BNFI (Bayesian NFI).
The posterior expectations (EAP), standard deviations, quantile-based credible intervals, and modes of these distributions are reported by the summary() method.
Example: three-factor CFA
We fit a three-factor CFA on the Holzinger–Swineford (1939) data.
Scalar fit measures
fitMeasures() returns the posterior mean (EAP) of each Bayesian fit index alongside the marginal log-likelihood, ppp, DIC, and gradient diagnostics.
fitMeasures(fit)
#> npar margloglik ppp dic p_dic BRMSEA
#> 21 -3823.429 0.000 7517.826 20.947 0.092
#> BGammaHat adjBGammaHat BMc
#> 0.957 0.919 0.904Posterior distributions of fit indices
To inspect the full posterior distribution of each index, use bfit_indices(). This returns an S3 object of class "bfit_indices".
bfi <- bfit_indices(fit)
bfi
#> Posterior summary of devM-based Bayesian fit indices (nsamp = 1000):
#>
#> BRMSEA BGammaHat adjBGammaHat BMc
#> 0.092 0.957 0.919 0.903Calling summary() provides a table of posterior summaries (Mean, SD, 2.5%, 50%, 97.5%, Mode) for each index.
summary(bfi)
#>
#> Posterior summary of devM-based Bayesian fit indices (nsamp = 1000):
#>
#> Mean SD X2.5. X25. X50. X75. X97.5. Mode
#> BRMSEA 0.092 0.005 0.084 0.088 0.092 0.095 0.103 0.091
#> BGammaHat 0.957 0.004 0.947 0.954 0.957 0.960 0.964 0.958
#> adjBGammaHat 0.919 0.008 0.900 0.914 0.920 0.925 0.932 0.921
#> BMc 0.903 0.010 0.881 0.897 0.904 0.910 0.919 0.905You can also access the raw per-sample vectors for custom analysis:
hist(bfi$indices$BRMSEA, breaks = 30, main = "BRMSEA", xlab = "BRMSEA",
col = "steelblue", border = "white")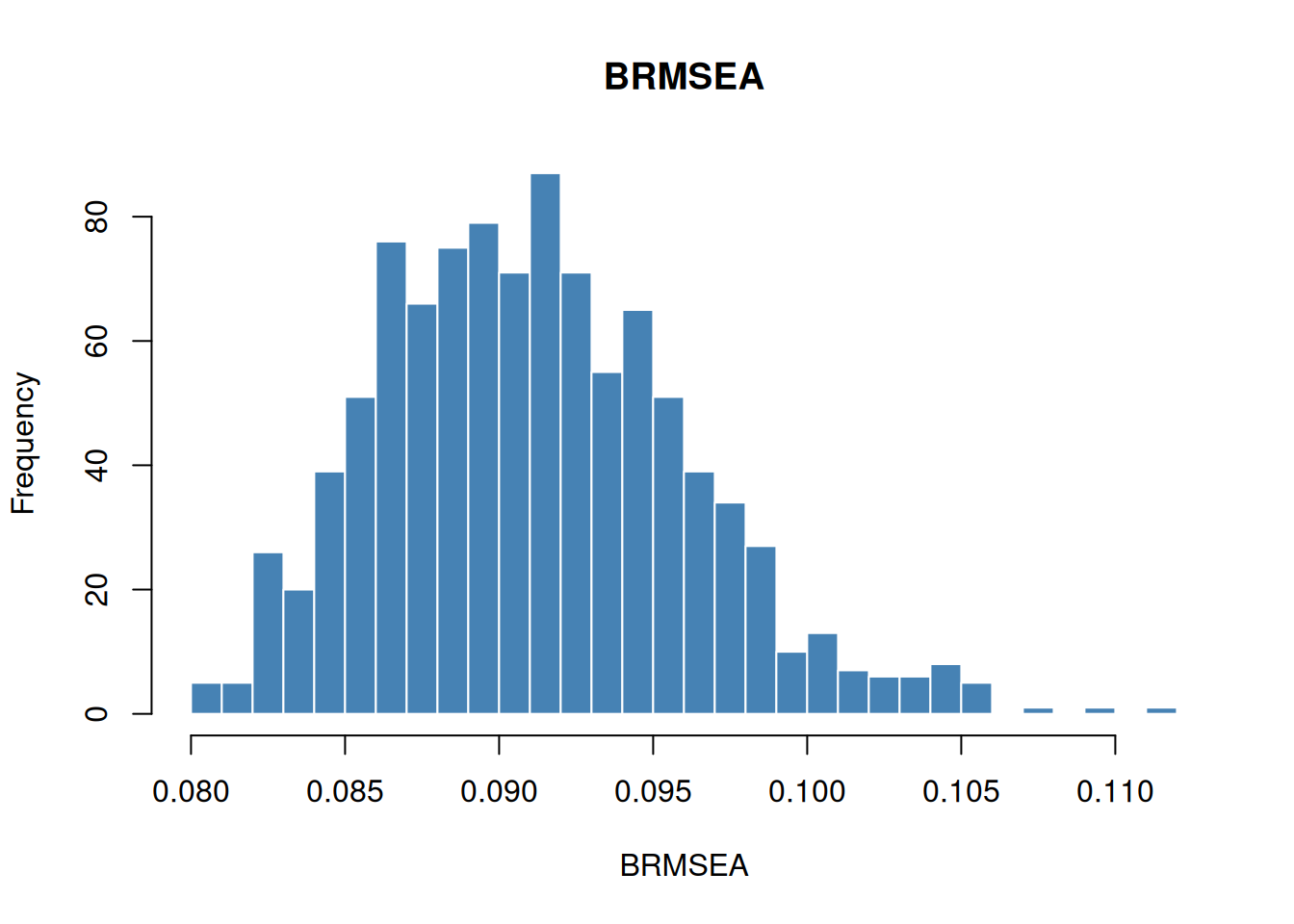
Comparison with a baseline model
Incremental indices (BCFI, BTLI, BNFI) require a baseline model. A common choice is the independence (uncorrelated) model.
null.model <- "
x1 ~~ x1
x2 ~~ x2
x3 ~~ x3
x4 ~~ x4
x5 ~~ x5
x6 ~~ x6
x7 ~~ x7
x8 ~~ x8
x9 ~~ x9
"
fit_null <- acfa(null.model, HolzingerSwineford1939, verbose = FALSE)Now pass the baseline model to fitMeasures() or bfit_indices():
fitMeasures(fit, baseline.model = fit_null)
#> npar margloglik ppp dic p_dic BRMSEA
#> 21 -3823.429 0.000 7517.826 20.947 0.092
#> BGammaHat adjBGammaHat BMc BCFI BTLI BNFI
#> 0.957 0.919 0.903 0.931 0.896 0.907
bfi_inc <- bfit_indices(fit, baseline.model = fit_null)
summary(bfi_inc)
#>
#> Posterior summary of devM-based Bayesian fit indices (nsamp = 1000):
#>
#> Mean SD X2.5. X25. X50. X75. X97.5. Mode
#> BRMSEA 0.092 0.005 0.084 0.088 0.092 0.095 0.103 0.089
#> BGammaHat 0.957 0.004 0.946 0.954 0.957 0.960 0.964 0.959
#> adjBGammaHat 0.919 0.008 0.900 0.914 0.920 0.925 0.932 0.924
#> BMc 0.903 0.010 0.880 0.897 0.904 0.910 0.918 0.909
#> BCFI 0.930 0.007 0.913 0.926 0.931 0.936 0.942 0.935
#> BTLI 0.896 0.011 0.870 0.889 0.897 0.904 0.913 0.903
#> BNFI 0.907 0.007 0.890 0.903 0.908 0.912 0.918 0.911Rescaling: "devM" vs "MCMC"
The rescale argument controls how the chi-square and degrees of freedom are computed. The default is "devM", which subtracts the DIC-based from the deviance. To use the classical scaling instead, set rescale = "MCMC":
bfi_mcmc <- bfit_indices(fit, rescale = "MCMC")
summary(bfi_mcmc)
#>
#> Posterior summary of MCMC-based Bayesian fit indices (nsamp = 1000):
#>
#> Mean SD X2.5. X25. X50. X75. X97.5. Mode
#> BRMSEA 0.107 0.004 0.100 0.104 0.106 0.110 0.116 0.105
#> BGammaHat 0.943 0.004 0.933 0.940 0.943 0.946 0.950 0.944
#> adjBGammaHat 0.892 0.008 0.874 0.887 0.893 0.898 0.905 0.895
#> BMc 0.872 0.010 0.850 0.866 0.873 0.879 0.887 0.875The two methods will generally produce different results, especially with informative priors or when deviates substantially from .
Differences from blavaan
INLAvaan’s Bayesian fit indices follow the same mathematical framework as blavaan (Merkle et al. 2021), with a few implementation differences:
| Feature | INLAvaan | blavaan |
|---|---|---|
| Posterior samples | INLA-based (Sobol/NORTA) | MCMC draws from Stan/JAGS |
| Rescaling methods |
"devM", "MCMC"
|
"devM", "MCMC", "ppmc"
|
| Effective parameters () | DIC-based only | DIC-based , LOOIC-based , or WAIC-based |
| HPD intervals | Not currently computed | Computed via {coda}
|
| Summary statistics | Mean, SD, 2.5%, 50%, 97.5%, Mode | EAP, Median, MAP, SD, HPD |
| Return class | S3 "bfit_indices"
|
S4 "blavFitIndices"
|
Currently, INLAvaan only supports from the DIC (i.e., p_dic). The "ppmc" rescaling method (which computes replicated data under the posterior predictive) is not yet available.