library(INLAvaan)
library(blavaan)
#> Loading required package: Rcpp
#> This is blavaan 0.5-10
#> On multicore systems, we suggest use of future::plan("multicore") or
#> future::plan("multisession") for faster post-MCMC computations.
set.seed(161)
# Generate data
n <- 250
truval <- c(0.8, 0.7, 0.6, 0.5, 0.4, -1.43, -0.55, -0.13, -0.72, -1.13)
dat <- lavaan::simulateData(
"eta =~ 0.8*y1 + 0.7*5y2 + 0.6*y3 + 0.5*y4 + 0.4*y5
y1 | -1.43*t1
y2 | -0.55*t1
y3 | -0.13*t1
y4 | -0.72*t1
y5 | -1.13*t1",
ordered = TRUE,
sample.nobs = n
)
head(dat)
#> y1 y2 y3 y4 y5
#> 1 2 2 1 2 2
#> 2 2 1 1 1 2
#> 3 2 2 2 1 2
#> 4 1 1 1 1 2
#> 5 2 2 1 2 2
#> 6 2 2 1 1 1
# Fit INLAvaan model
mod <- "eta =~ y1 + y2 + y3 + y4 + y5"
fit <- acfa(mod, dat, ordered = TRUE, std.lv = TRUE, estimator = "PML")
#> ℹ Finding posterior mode.
#> ✔ Finding posterior mode. [150ms]
#>
#> ℹ Computing the Hessian.
#> ✔ Computing the Hessian. [85ms]
#>
#> ℹ Performing VB correction.
#> ✔ VB correction; mean |δ| = 0.403σ. [408ms]
#>
#> ⠙ Fitting 0/10 skew-normal marginals.
#> ✔ Fitting 10/10 skew-normal marginals. [653ms]
#>
#> ℹ Adjusting copula correlations (NORTA).
#> ✔ Adjusting copula correlations (NORTA). [44ms]
#>
#> ⠙ Posterior sampling and summarising.
#> ⠹ Posterior sampling and summarising.
#> ✔ Posterior sampling and summarising. [896ms]
#>
summary(fit)
#> INLAvaan 0.2.4.9001 ended normally after 37 iterations
#>
#> Estimator BAYES
#> Optimization method NLMINB
#> Number of model parameters 10
#>
#> Number of observations 250
#>
#> Model Test (User Model):
#>
#> Marginal log-likelihood -1094.328
#> PPP (Chi-square) 0.000
#>
#> Information Criteria:
#>
#> Deviance (DIC) 2126.012
#> Effective parameters (pD) 7.420
#>
#> Parameter Estimates:
#>
#> Parameterization Theta
#> Marginalisation method SKEWNORM
#> VB correction TRUE
#>
#> Latent Variables:
#> Estimate SD 2.5% 97.5% NMAD Prior
#> eta =~
#> y1 0.825 0.329 0.302 1.576 0.049 normal(0,10)
#> y2 0.640 0.248 0.244 1.205 0.025 normal(0,10)
#> y3 0.627 0.237 0.235 1.157 0.011 normal(0,10)
#> y4 1.121 0.442 0.473 2.154 0.013 normal(0,10)
#> y5 0.520 0.225 0.149 1.025 0.017 normal(0,10)
#>
#> Thresholds:
#> Estimate SD 2.5% 97.5% NMAD Prior
#> y1|t1 -2.195 0.353 -3.018 -1.670 0.030 normal(0,1.5)
#> y2|t1 -0.795 0.127 -1.087 -0.600 0.061 normal(0,1.5)
#> y3|t1 -0.300 0.083 -0.482 -0.155 0.013 normal(0,1.5)
#> y4|t1 -0.994 0.222 -1.514 -0.673 0.034 normal(0,1.5)
#> y5|t1 -1.239 0.153 -1.592 -1.006 0.072 normal(0,1.5)
#>
#> Variances:
#> Estimate SD 2.5% 97.5% NMAD Prior
#> .y1 1.000
#> .y2 1.000
#> .y3 1.000
#> .y4 1.000
#> .y5 1.000
#> eta 1.000
#>
#> Scales y*:
#> Estimate SD 2.5% 97.5% NMAD Prior
#> y1 1.325
#> y2 1.203
#> y3 1.194
#> y4 1.519
#> y5 1.141
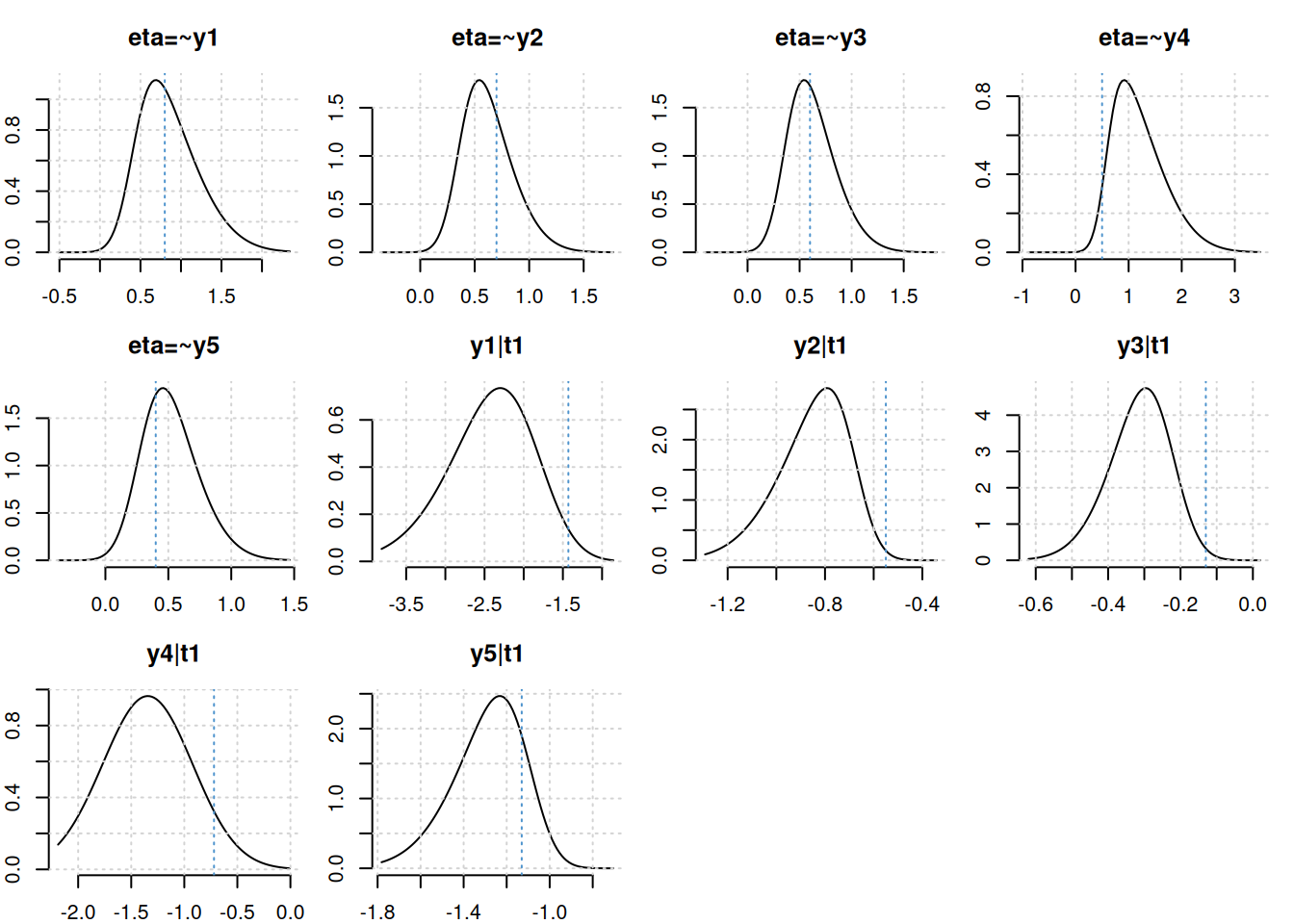
plot(fit, truth = truval)