library(INLAvaan)
mod <- "
# Latent variable definitions
ind60 =~ x1 + x2 + x3
dem60 =~ y1 + y2 + y3 + y4
dem65 =~ y5 + y6 + y7 + y8
# Latent regressions
dem60 ~ ind60
dem65 ~ ind60 + dem60
# Residual correlations
y1 ~~ y5
y2 ~~ y4 + y6
y3 ~~ y7
y4 ~~ y8
y6 ~~ y8
"
dat <- lavaan::PoliticalDemocracy
# Simulate missingness (MCAR)
set.seed(221)
mis <- matrix(rbinom(prod(dim(dat)), 1, 0.99), nrow(dat), ncol(dat))
datmiss <- dat * mis
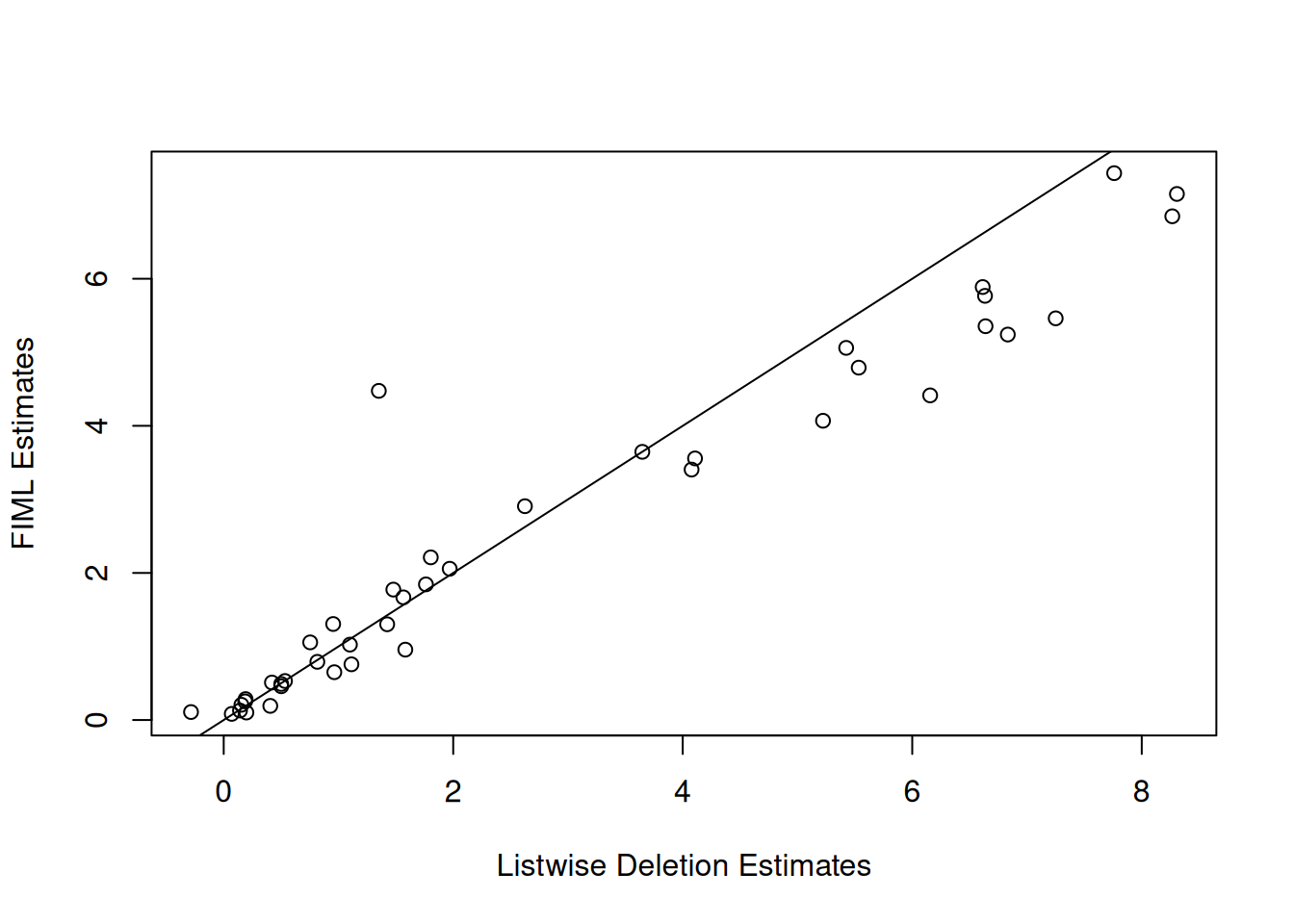
datmiss[datmiss == 0] <- NAINLAvaan handles missing data in one of two ways: listwise deletion (default, i.e. uses all complete cases) or Full Information Maximum Likelihood (FIML; missing = "ML").
Simulate Data
Listwise Deletion
fit1 <- asem(mod, datmiss, meanstructure = TRUE)
#> ℹ Finding posterior mode.
#> ✔ Finding posterior mode. [132ms]
#>
#> ℹ Computing the Hessian.
#> ✔ Computing the Hessian. [147ms]
#>
#> ℹ Performing VB correction.
#> ✔ VB correction; mean |δ| = 0.222σ. [458ms]
#>
#> ⠙ Fitting 0/42 skew-normal marginals.
#> ⠹ Fitting 25/42 skew-normal marginals.
#> ✔ Fitting 42/42 skew-normal marginals. [3s]
#>
#> ℹ Adjusting copula correlations (NORTA).
#> ✔ Adjusting copula correlations (NORTA). [341ms]
#>
#> ⠙ Posterior sampling and summarising.
#> ✔ Posterior sampling and summarising. [797ms]
#>
fit1@Data@nobs[[1]] == nrow(datmiss[complete.cases(datmiss), ])
#> [1] TRUE
print(fit1)
#> INLAvaan 0.2.4.9001 ended normally after 71 iterations
#>
#> Estimator BAYES
#> Optimization method NLMINB
#> Number of model parameters 42
#>
#> Used Total
#> Number of observations 35 75
#>
#> Model Test (User Model):
#>
#> Marginal log-likelihood -818.478
#> PPP (Chi-square) 0.499
coef(fit1)
#> ind60=~x2 ind60=~x3 dem60=~y2 dem60=~y3 dem60=~y4 dem65=~y6
#> 1.808 1.750 0.943 0.805 1.472 1.061
#> dem65=~y7 dem65=~y8 dem60~ind60 dem65~ind60 dem65~dem60 y1~~y5
#> 0.721 1.354 0.913 0.567 1.080 0.469
#> y2~~y4 y2~~y6 y3~~y7 y4~~y8 y6~~y8 x1~~x1
#> 1.844 3.596 0.607 -0.669 1.283 0.071
#> x2~~x2 x3~~x3 y1~~y1 y2~~y2 y3~~y3 y4~~y4
#> 0.144 0.424 1.698 7.790 4.086 2.786
#> y5~~y5 y6~~y6 y7~~y7 y8~~y8 ind60~~ind60 dem60~~dem60
#> 1.537 6.752 2.038 3.983 0.506 1.439
#> dem65~~dem65 x1~1 x2~1 x3~1 y1~1 y2~1
#> 0.139 5.424 5.533 4.107 7.250 6.634
#> y3~1 y4~1 y5~1 y6~1 y7~1 y8~1
#> 8.306 6.833 6.639 5.223 8.266 6.156Full Information Maximum Likelihood (FIML)
fit2 <- asem(mod, datmiss, missing = "ML", meanstructure = TRUE)
#> ℹ Finding posterior mode.
#> ✔ Finding posterior mode. [247ms]
#>
#> ℹ Computing the Hessian.
#> ✔ Computing the Hessian. [234ms]
#>
#> ℹ Performing VB correction.
#> ✔ VB correction; mean |δ| = 0.160σ. [490ms]
#>
#> ⠙ Fitting 0/42 skew-normal marginals.
#> ⠹ Fitting 19/42 skew-normal marginals.
#> ✔ Fitting 42/42 skew-normal marginals. [5.2s]
#>
#> ℹ Adjusting copula correlations (NORTA).
#> ✔ Adjusting copula correlations (NORTA). [406ms]
#>
#> ⠙ Posterior sampling and summarising.
#> ✔ Posterior sampling and summarising. [1.3s]
#>
print(fit2)
#> INLAvaan 0.2.4.9001 ended normally after 93 iterations
#>
#> Estimator BAYES
#> Optimization method NLMINB
#> Number of model parameters 42
#>
#> Number of observations 75
#> Number of missing patterns 19
#>
#> Model Test (User Model):
#>
#> Marginal log-likelihood -1414.868
#> PPP (Chi-square) 0.005
coef(fit2)
#> ind60=~x2 ind60=~x3 dem60=~y2 dem60=~y3 dem60=~y4 dem65=~y6
#> 2.214 1.835 0.650 0.791 0.954 1.026
#> dem65=~y7 dem65=~y8 dem60~ind60 dem65~ind60 dem65~dem60 y1~~y5
#> 1.050 1.287 1.299 0.524 0.756 0.482
#> y2~~y4 y2~~y6 y3~~y7 y4~~y8 y6~~y8 x1~~x1
#> 0.898 3.316 0.301 0.365 1.294 0.084
#> x2~~x2 x3~~x3 y1~~y1 y2~~y2 y3~~y3 y4~~y4
#> 0.136 0.511 1.699 7.448 3.408 2.945
#> y5~~y5 y6~~y6 y7~~y7 y8~~y8 ind60~~ind60 dem60~~dem60
#> 1.794 5.910 2.073 3.682 0.462 4.558
#> dem65~~dem65 x1~1 x2~1 x3~1 y1~1 y2~1
#> 0.206 5.060 4.791 3.557 5.462 5.787
#> y3~1 y4~1 y5~1 y6~1 y7~1 y8~1
#> 7.158 5.253 5.356 4.092 6.855 4.422